- 首页
- 科研与创新
科研与创新

精选科学刊物
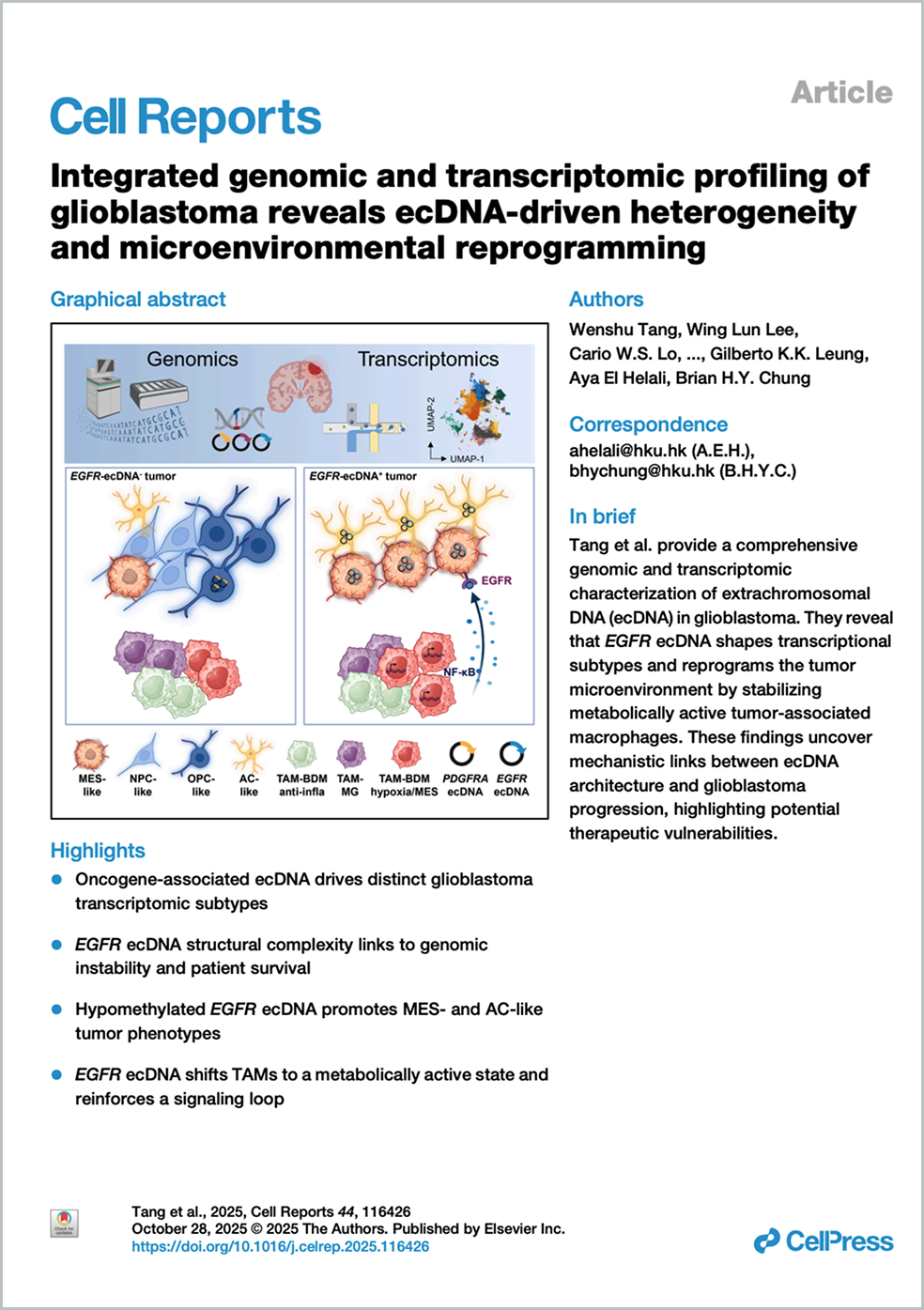
Integrated genomic and transcriptomic profiling of glioblastoma reveals ecDNA-driven heterogeneity and microenvironmental reprogramming (请参阅英文版本)
Glioblastoma (GB) is an aggressive brain tumor with limited treatment options, making it crucial to integrate genomic and transcriptomic profiling to identify genetic alterations and cellular functional states. We perform short- and long-read whole-genome sequencing (WGS) and single-nucleus RNA sequencing on 42 Chinese GB patients to characterize the intra- and extrachromosomal mutation landscape, cell-type composition, and pathway activity. Our analysis identifies amplified oncogenes, including EGFR, MYC, CDK4, PDGFRA, and PPARGC1A, localized on extrachromosomal DNA (ecDNA). Notably, EGFR ecDNA harbors distinct structures that correlate with patient survival and exhibit a unique DNA methylation pattern that influences gene expression, driving malignant cell differentiation toward MES-like and AC-like subtypes. Specifically, EGFR ecDNA stabilizes tumor-associated macrophages in a hypoxia- and metabolism-driven state, reinforcing a reciprocal AREG-EGFR signaling loop with mesenchymal-like tumor cells. Together, these findings uncover a mechanistic link between ecDNA architecture, transcriptional subtypes, and microenvironmental remodeling, offering critical insights for advancing precision oncology in GB.

Identification of technically challenging variants: Whole-genome sequencing improves diagnostic yield in patients with high clinical suspicion of rare diseases (请参阅英文版本)
The total burden of rare diseases is significant worldwide, with over 300 million people being affected. Many rare diseases have both well-defined clinical phenotypes and established genetic causes. However, a remarkable proportion of patients with high clinical suspicion of a rare disease remain genetically undiagnosed and stuck in the diagnostic odyssey after having a cascade of conventional genetic tests. One of the major factors contributing to this is that many types of variants are technically intractable to whole-exome sequencing (WES). In this study, the added diagnostic power of whole-genome sequencing (WGS) for patients with clinically suspected rare diseases was assessed by detecting technically challenging variants. 3,169 patients from the Hong Kong Genome Project (HKGP) were reviewed, identifying 322 individuals having high clinical suspicion of a rare disorder with well-established genetic etiology. Notably, 180 patients have performed at least one previous genetic test. Through PCR-free short-read WGS and a comprehensive in-house analytic pipeline, causative variants were found in 138 patients (138 of 322, 42.9%), 30 of which (30 of 138, 21.7%) are attributed to technically challenging variants. These included 6 variants in low-coverage regions with PCR bias, 2 deep intronic variants, 2 repeat expansions, 19 structural variants, and 2 variants in genes with a homologous pseudogene. The study demonstrated the indispensable diagnostic power of WGS in detecting technically challenging variants and the capability to serve as an all-in-one test for patients with high clinical suspicion of rare diseases.

The implementation of genome sequencing in rare genetic diseases diagnosis: a pilot study from the Hong Kong genome project (请参阅英文版本)
Genome sequencing (GS) has revolutionised the diagnostic odyssey of patients with rare genetic diseases (RDs) and accelerated large-scale genome projects globally. However, the impact of GS on patients with RDs is yet to be investigated among genome projects in Asia. The Hong Kong Genome Project (HKGP) was implemented to benefit patients and families with RDs in Hong Kong, and to increase the inclusiveness of Chinese genomic data. This study evaluated the impact of short read GS (srGS), complemented by long read GS (lrGS) in a subset, on individuals recruited in the pilot phase of the HKGP.

Meta-analysis of the diagnostic and clinical utility of exome and genome sequencing in pediatric and adult patients with rare diseases across diverse populations (请参阅英文版本)
Purpose:
This meta-analysis aims to compare the diagnostic and clinical utility of exome sequencing (ES) vs genome sequencing (GS) in pediatric and adult patients with rare diseases across diverse populations.
Methods:
A meta-analysis was conducted to identify studies from 2011 to 2021.
Results:
One hundred sixty-one studies across 31 countries/regions were eligible, featuring 50,417 probands of diverse populations. Diagnostic rates of ES (0.38, 95% CI 0.36-0.40) and GS (0.34, 95% CI 0.30-0.38) were similar (P = .1). Within-cohort comparison illustrated 1.2-times odds of diagnosis by GS over ES (95% CI 0.79-1.83, P = .38). GS studies discovered a higher range of novel genes than ES studies; yet, the rate of variant of unknown significance did not differ (P = .78). Among high-quality studies, clinical utility of GS (0.77, 95% CI 0.64-0.90) was higher than that of ES (0.44, 95% CI 0.30-0.58) (P < .01).
Conclusion:
This meta-analysis provides an important update to demonstrate the similar diagnostic rates between ES and GS and the higher clinical utility of GS over ES. With the newly published recommendations for clinical interpretation of variants found in noncoding regions of the genome and the trend of decreasing variant of unknown significance and GS cost, it is expected that GS will be more widely used in clinical settings.






.png)
.png)
.png)
.png)